Histone Modifications in Acute Myeloid Leukemia
In collaboration with:
Description: Histone post-translational modifications (PTMs) create a powerful regulatory mechanism for maintaining chromosomal integrity in cells. Histone acetylation and methylation, the most widely studied histone PTMs, act in concert with chromatin-associated proteins to control access to genetic information during transcription. Alterations in cellular histone PTMs have been linked to disease states and have crucial biomarker and therapeutic potential. Traditional bottom-up mass spectrometry of histones requires large numbers of cells, typically one million or more. Decreasing the required number of cells is pivotal for the success of projects with low input amounts.
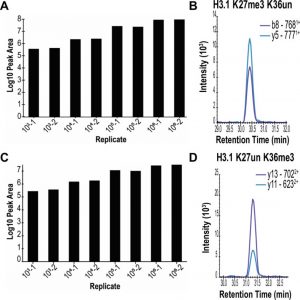
Lowering the input cell amount for assessment of histone marks. Peak area comparison of representative histone marks generated from 103−106 HeLa-S3 cells for (A) H3.1 K27un K36me3 and (C) H3.1 K27me3 K36un and representative chromatograms (B) and (D), respectively.
Project status: Completed
Publications:
-Histone H1 loss drives lymphoma by disrupting 3D chromatin architecture. Nevin Yusufova et al.
-Mutant EZH2 Induces a Pre-malignant Lymphoma Niche by Reprogramming the Immune Response. Wendy Beguelin et al.
-Targeted detection and quantitation of histone modifications from 1,000 cells. Nebiyu A. Abshiru et al.
Congress Presentations:
– ASMS 2017
Spinoff funding: none





Comments are closed.