21 st Century Analysis of Histone Modifications
Overview
It is well-known that distinct histone modification states carry key information regarding gene transcription. Histone methylation, in particular, can transmit information through mitosis, forming the basis of an epiproteomic “fingerprint.” In TR&D 4, we will establish and validate a full pipeline to quantitate all major sites of lysine methylation from 10
6
down to 10
3
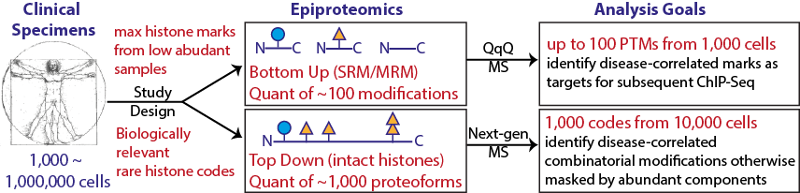
cells. The output of this pipeline will be a quantitative report of modification levels spanning 5 orders of magnitude in abundance for >100 discrete modification states. To accomplish this feat, we propose a bifurcated approach using two different mass spectrometric strategies for protein analysis. The platform will include interrogating full-length histones (top-down) and triple-quadrupole based tryptic peptide quantitation (bottom-up).
Top Down Analysis of Histone H3
Histone H3 is the most highly modified core histone protein. In a
recent publication in
Molecular and Cellular Proteomics
, we brought together the collaborating lab of Dr. Jonathan Licht with our instrumentation and data analysis teams to explore the different histone proteoforms present in a multiple myeloma cell line. The result was a deep and thorough analysis of nearly 100 different proteoforms of histone H3.1 and H3.3 where we recapitulated the antagonistic crosstalk between K27 and K36 methylation in H3.1.


Comments are closed.